PLANT ORGANELLE GENE EDITING PLATFORM
GREENedit™
The GREENedit™ platform is a world-first, exclusive technology enabling precise, targeted gene editing in both chloroplasts and mitochondria. Leveraging extremely high transcription rates and rapid attainment of homoplasmy, it allows the accurate and efficient generation of versatile plant models for a wide range of applications.
By directly engineering organelle genomes, GREENedit™ overcomes the limitations of nuclear editing, unlocking unprecedented control over photosynthesis, energy metabolism, and other key cellular pathways. This platform sets a new standard for plant biotechnology, combining precision, speed, and scalability. Its robust and flexible design makes it an ideal tool for advanced research and commercial plant engineering.
WHY CHLOROPLASTS?
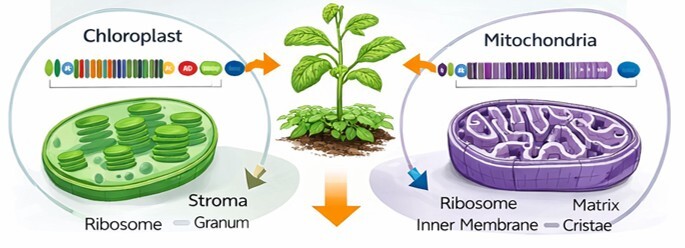
Chloroplasts are the powerhouses behind photosynthesis, helping plants grow and thrive. By editing chloroplast genes, we can directly boost photosynthesis and carbon fixation, making plants more productive. Chloroplasts carry many copies of their DNA, so changes spread quickly and reliably. They also have their own machinery for making proteins, which means high-level, stable expression. Plus, targeting chloroplasts avoids unintended effects in the plant’s nuclear DNA. In short, it’s a smart, safe way to make plants stronger and more efficient.
Mitochondria are the energy factories of plant cells, keeping everything running smoothly. Editing mitochondrial genes lets us improve energy use, growth, and stress resilience right at the cellular level. Like chloroplasts, mitochondria have multiple DNA copies, so edits take hold effectively. Nuclear editing can’t reach mitochondria directly, which makes them a unique opportunity for innovation. With precise changes, we can help plants grow better, handle stress, and adapt to new environments. It’s a powerful way to unlock the hidden potential inside every plant cell.
WHY MITOCHONDRIA?
GENERATING VALUABLE VARIATION
THROUGH PRECISION-TARGETED EDITS
PLANT ORGANELLE GENE EDITING PROCESS
Plant organelle gene editing involves precise modification of the genomes within chloroplasts or mitochondria, typically achieved through delivery of engineered nucleases or gene-editing tools into the organelles. This process requires overcoming challenges such as targeting multiple genome copies to achieve homoplasmy and utilizing organelle-specific expression systems. Successful editing enables the alteration of key metabolic pathways to enhance plant traits like photosynthesis, stress tolerance, and energy efficiency.
GENOME CONSTRUCT DESIGN & CLONING


IN-VIVO ORGANELLE GENE EDITING
MODEL PLANT (Arabidopsis) VALIDATION

Trait Transfer into various plants through
PLANT REGENERATION
TECHNOLOGY HIGHLIGHTS

Organelle-specific Targeting
Precise subcellular targeting
to chloroplasts and mitochondria

Photosynthesis Optimization Library
Offers extensive editing database

Minimized off-target effects
Significantly limits unintended edits at Non-Target sites and Bystanders

AI-Driven Gene Editing Design
Fast and flexible gene editing design powered by AI
RESEARCH PUBLICATIONS
Herbicide-resistant plants produced by precision adenine base editing in plastid DNA
Nature Plants, 2024, 10, 1652-1658
Mok YG, Hong SH, Seo DI, Choi SH, Kim HK, Jin DM, Lee JE, Kim JS
Prime Editing with genuine Cas9 nickases minimizes unwanted indels
Nature Communications, 2023, 14, Article 1786
Lee JS, Lim KY, Kim A, Mok YG, Chung EG, Cho SI, Lee JM, Kim JS
Targeted A-to-G base editing in human mitochondrial DNA with programmable deaminases
Cell, 2022, 185, 1764-1776
Cho SI, Lee S, Mok YG, Lim K, Lee J, Lee JM, Chung E, Kim JS
Base editing in human cells with monomeric DddA-TALE fusion deaminases
Nature Communications, 2022, 13, Article 4038
Mok YG, Lee JM, Chung E, Lee J, Lim K, Cho SI, Kim JS
Targeted A-to-G base editing of chloroplast DNA in plants
Nature Plants, 2022, 8, 1378-1384
Mok YG, Hong S, Bae SJ, Cho SI, Kim JS
Chloroplast and mitochondrial DNA editing in plants
Nature Plants, 2021, 7, 899-905
Kang BC, Bae SJ, Lee S, Lee JS, Kim A, Lee H, Baek G, Seo H, Kim J, Kim JS
